Getting Started¶
Here, we exemplify the basic usage of ParallelKDEpy with the core estimator gradepro. For more details about the existing estimators, their parameters, as well as the more in-depth implementation details, please refer to the ParallelKDE.jl documentation.
To estimate a density on CPU with a default grid using the gradepro method, you can use the following code:
import numpy as np
import matplotlib.pyplot as plt
import parallelkdepy as pkde
data = np.random.normal(size=(10000, 1)) # Example data: 10000 samples in 1D
density_estimation = pkde.DensityEstimation(
data,
grid=True,
device="cpu",
)
density_estimation.estimate_density("gradepro")
density_estimated = density_estimation.get_density()
grid_coordinates = density_estimation.generate_grid().to_meshgrid()[0]
# Evaluate true density for comparison
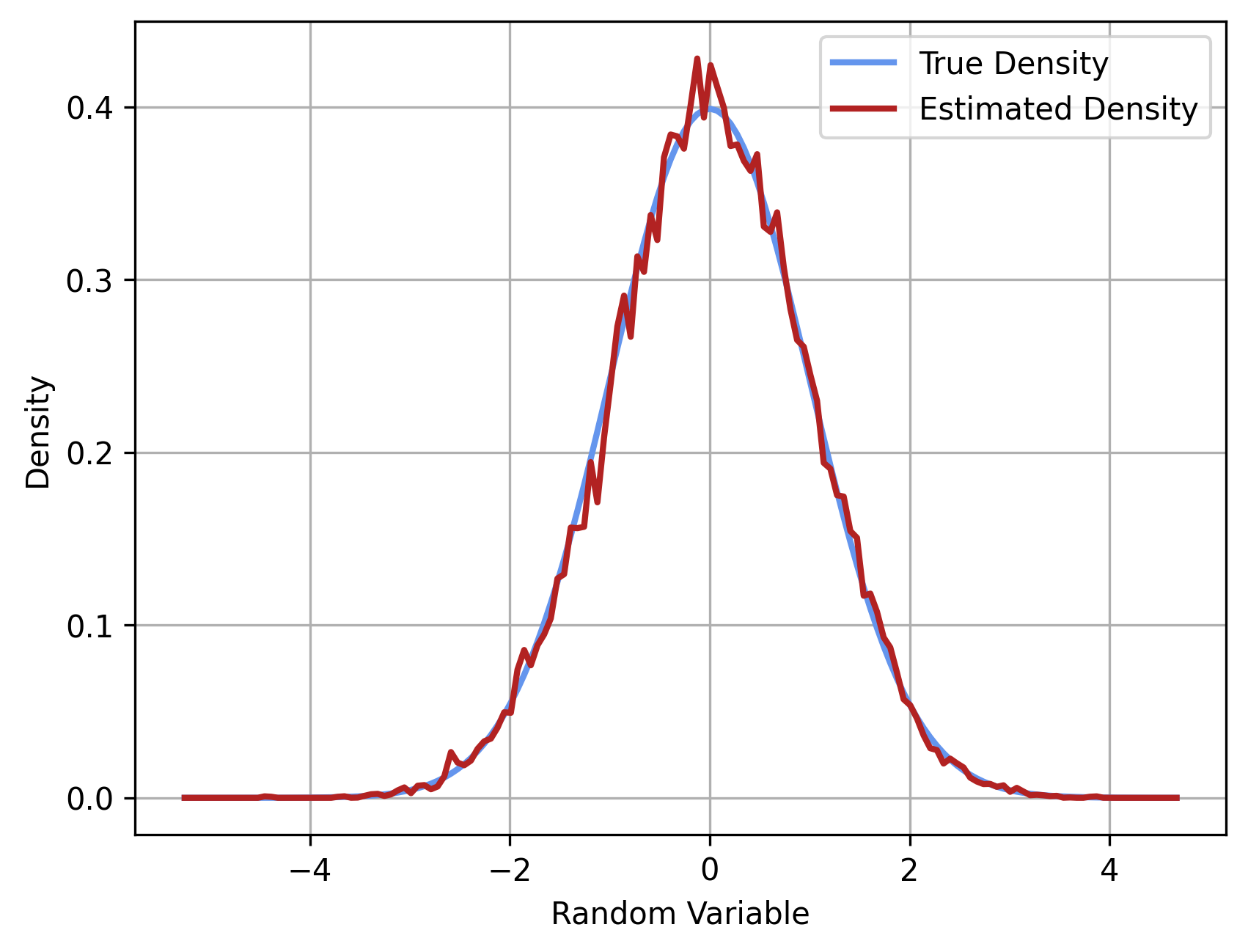
density_true = np.exp(-0.5 * grid_coordinates**2) / np.sqrt(2 * np.pi)
plt.plot(grid_coordinates, density_true, label="True Density", lw=2, c="cornflowerblue")
plt.plot(
grid_coordinates, density_estimated, label="Estimated Density", lw=2, c="firebrick"
)
plt.xlabel("Random Variable")
plt.ylabel("Density")
plt.legend()
plt.grid()
That’s it! See the API Reference for more details on the available methods and parameters.